CRISPR Associated Transposons (CASTs)
The CRISPR Associated Transposon system allows for the targeted insertion of a large genetic cargo (up to 10 kb) into a bacterial genome roughly 50 bp downstream of where a 32 bp spacer sequence matches that genome. The system described here is based off of the Type I-F originally from Vibrio cholerae and can be used for a variety of applications. To become familiar with CRISPR Associated Transposons, it is recommended that you read the following papers as a primer: Hu et al. 2023, Klompe et al. 2019, Vo et al. 2021. Our lab uses the Ultra CAST system developed by Anthony. See the details of this system in https://pmc.ncbi.nlm.nih.gov/articles/PMC12358020/. The CAST system is encoded on the following plasmid which is Golden Gate compatible and contains a two dropouts: a GFP dropout with BbsI cut sites for the insertion of a spacer sequence and a mScarlet dropout with both BsmB1 (compatible with ConRE and ConLS overhangs) and BsaI (compatible with type 2-4 overhangs) cut sites.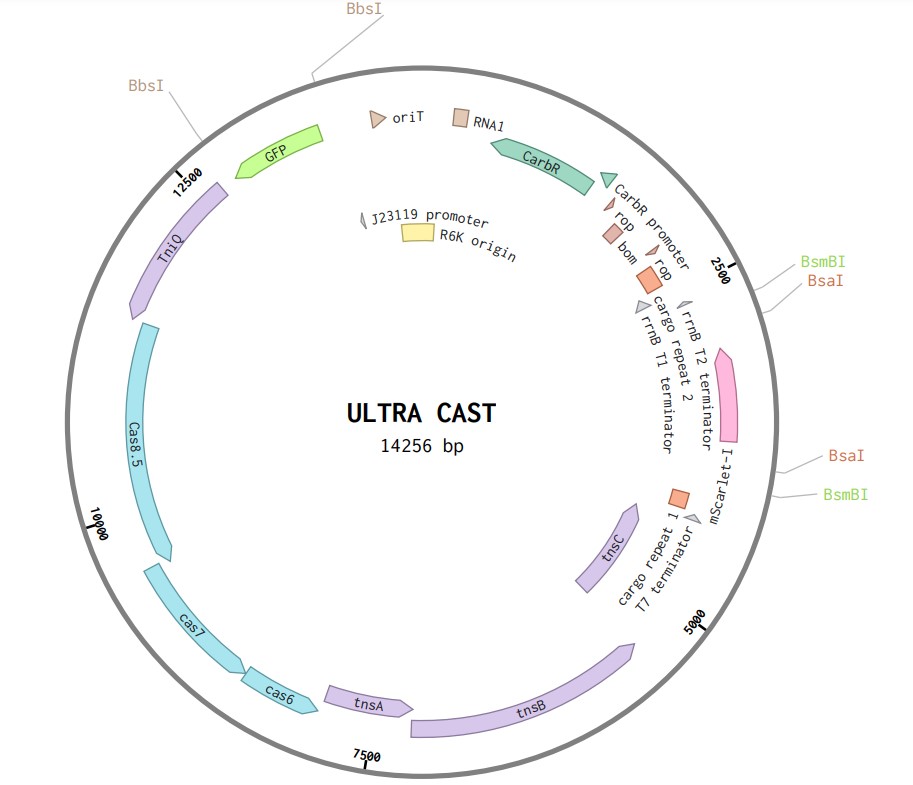 To design a spacer sequence, scan the genome for the PAM sequence 5'-CC-3'. Next, copy the preceding 32 bp sequence and add the following sequences:
At the 5' end of the spacer: GCATGAAGACAAaaactgaactgccgagtaggtagctgataac. This the upstream Bbs1 site and the CRIPSR repeat.
At the 3' end of the spacer: gtgaactgccgagtaggtagctgataaatggGTGTCTTCGCAT. This is the CRIPSR repeat and the downstream Bbs1 site
Keep in mind when designing the spacer sequence that the cargo will insert 50 bp DOWNSTREAM from the end of where the spacer sequence matches the genome.
The best way to order this is to order two ultramers. The first ultramer should include the upstream Bbs1 site, the 5' CRIPSR repeat and the 32bp gRNA sequence. The second ultramer should be the reverse complement of the downstream Bbs1 site, the CRISPR repeat and the 32-bp gRNA sequence. This way, the two ultramers will overlap over the 32bp gRNA sequence. Make sure you correctly use the reverse complement to anneal the ultramers!
To anneal the ultramers, resuspend in water to standard concentration of 10uM. Then add 10 uL of each ultramer to 80 uL of duplex buffer.
Duplex buffer:
To design a spacer sequence, scan the genome for the PAM sequence 5'-CC-3'. Next, copy the preceding 32 bp sequence and add the following sequences:
At the 5' end of the spacer: GCATGAAGACAAaaactgaactgccgagtaggtagctgataac. This the upstream Bbs1 site and the CRIPSR repeat.
At the 3' end of the spacer: gtgaactgccgagtaggtagctgataaatggGTGTCTTCGCAT. This is the CRIPSR repeat and the downstream Bbs1 site
Keep in mind when designing the spacer sequence that the cargo will insert 50 bp DOWNSTREAM from the end of where the spacer sequence matches the genome.
The best way to order this is to order two ultramers. The first ultramer should include the upstream Bbs1 site, the 5' CRIPSR repeat and the 32bp gRNA sequence. The second ultramer should be the reverse complement of the downstream Bbs1 site, the CRISPR repeat and the 32-bp gRNA sequence. This way, the two ultramers will overlap over the 32bp gRNA sequence. Make sure you correctly use the reverse complement to anneal the ultramers!
To anneal the ultramers, resuspend in water to standard concentration of 10uM. Then add 10 uL of each ultramer to 80 uL of duplex buffer.
Duplex buffer:
- 30 mM HEPES
- 100 mM potassium acetate
- pH 7.5
- 95C -> 2min
- 94C -> 30s
- 94C -> 12C in 30s intervals drop -1C
- 20 uL of annealed oligos from above
- 5 uL of NEB buffer 2
- 1 uL of dNTPs
- 1 uL of Klenow polymerase
- 23 uL of ddH2O
| I | Attachment | History | Action | Size | Date | Who | Comment |
|---|---|---|---|---|---|---|---|
| |
CAST_pic.jpg | r1 | manage | 70.5 K | 2024-04-09 - 21:47 | AnthonyVandieren | CAST_pic |
Barrick Lab > ProtocolList > ProtocolsCRISPRAssociatedTransposons(CASTs)

