Evolving and Engineering Insect Symbionts
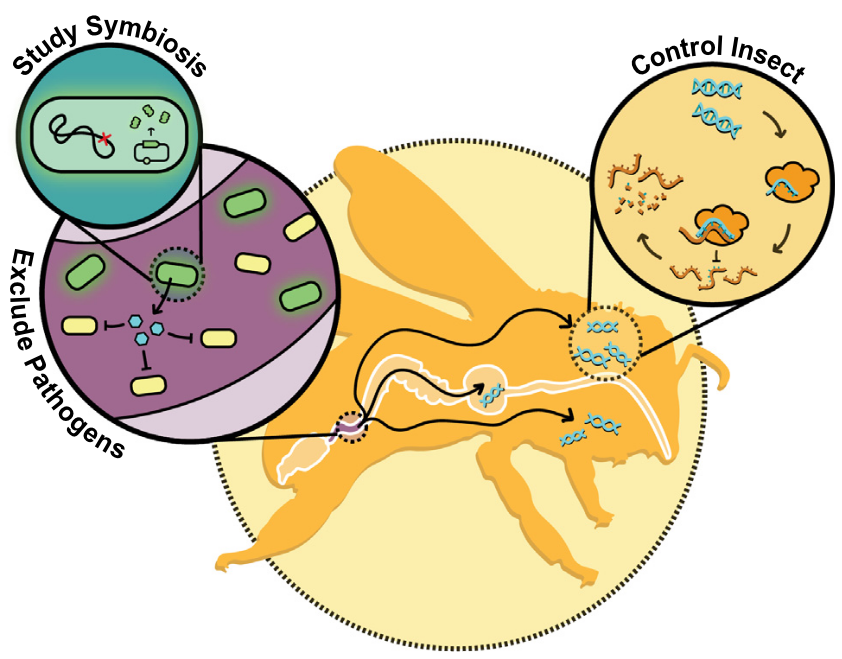
Insect symbiont engineering overview
Many insects have more consequential associations with bacteria than we have with the human microbiome. We create genetic tools to study these bacterial symbionts and engineer their interactions with insects. We also use experimental evolution to examine how new symbionts arise and how interactions within a microbiome and between microbes and their hosts can change. These projects are highly collaborative and have involved researchers from the
Moran Lab, the
Davies Lab, the
Ellington Lab and other research groups.
 Current Funding:
Current Funding: ARO MURI, NSF EDGE, USDA NIFA
 Past Funding:
Past Funding: DARPA BRICS, DARPA Insect Allies, ERDC
 Overview Publication
Overview Publication Honey Bees: Functional Genomics and Protecting Pollinator Health
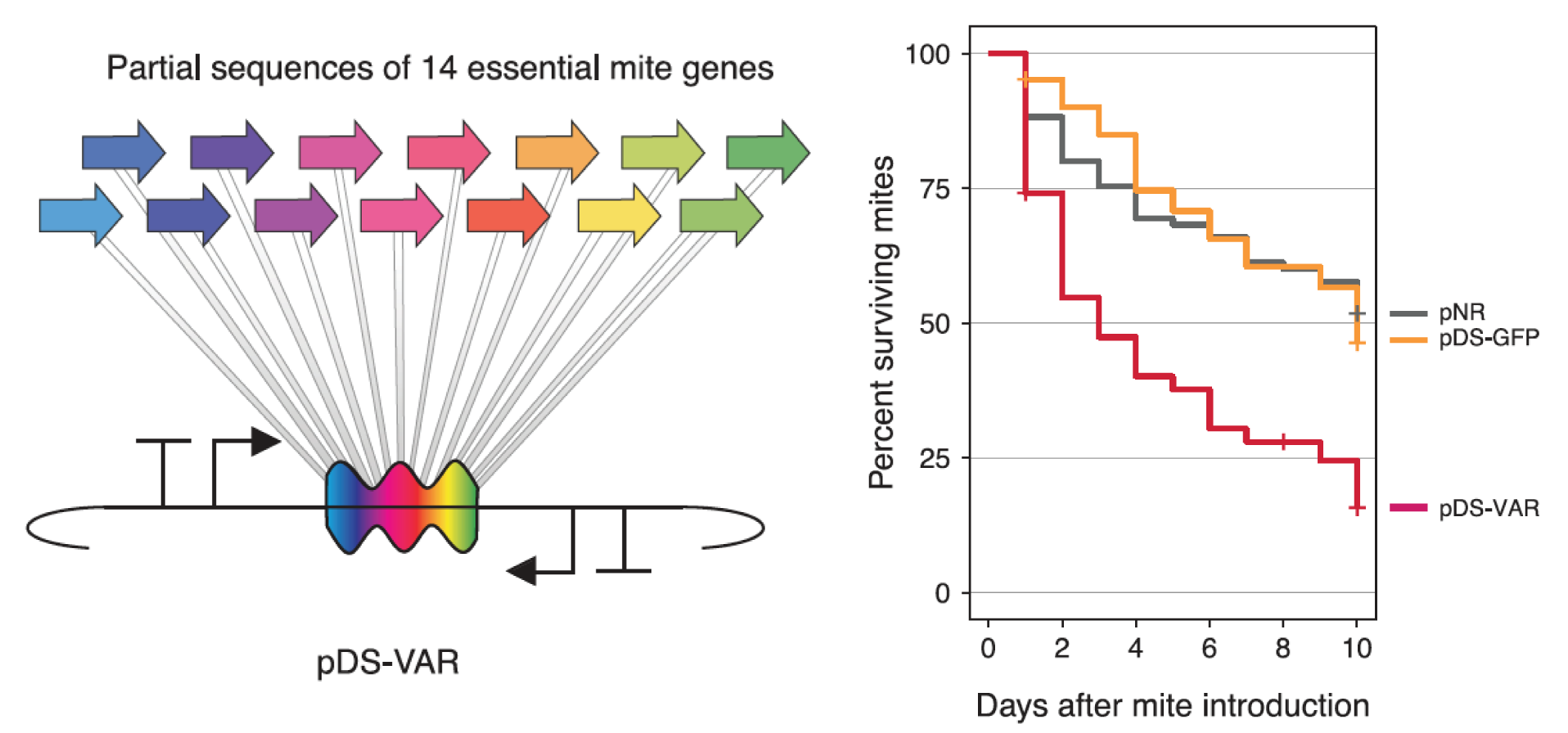
Engineered symbionts kill honey bee parasites
Bees are ecologically and economically important pollinators. We created a bee microbiome toolkit (BTK) for engineering bacteria that colonize the guts of honey bees (
Apis mellifera) and bumble bees (
Bombus species). We recently showed that we can engineer the bacterium
Snodgrassella alvi to implement effective symbiont-mediated RNAi. This system can be used for the targeted knockdown of bee genes to study their functions. For example, we have used it to alter honey bee feeding behavior. We have also shown that these engineered symbionts can be used to protect honey bees from viral pathogens and arthropod parasites that imperil the health of hives.
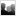 Current Researchers:
Current Researchers: Zuberi Ashraf, Kadena Cope, Korin Jones, Kathleen Sotelo, PJ Lariviere, Lucio Navarro, Dennis Mishler
 News: Bacteria Engineered to Protect Bees from Pests and Pathogens
News: Bacteria Engineered to Protect Bees from Pests and Pathogens
 Representative Publications
Representative Publications Aphids: Understanding the Evolution of Endosymbionts and Pest Control
GFP-tagged symbiont colonizing the aphid gut
Aphids are plant pests and model systems for studying bacteria-insect symbioses. We have developed tools for engineering strains of
Serratia symbiotica that colonize the aphid gut. We are now using these tools to unravel how these strains, which can be pathogenic to their hosts, are related to strains of
S. symbiotica that live within insect cells, are inherited across aphid generations, and can benefit their aphid hosts. We have showed that cultured
S. symbiotica strains are capable of maternal transmission within aphids, suggesting that they possess a latent capacity to evolve a long-term symbiotic relationship with their hosts. We are also exploring applications of these engineered bacteria related to pest control, especially through symbiont-mediated RNAi.
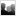 Current Researchers:
Current Researchers: Anthony VanDieren, Lucio Navarro
 News: Turning Plant Pests into Helpers
News: Turning Plant Pests into Helpers
 Representative Publications
Representative Publications
Improving the Reliability of Synthetic Biology
Cells that evolve mutations in a burdensome function can rapidly take over a population
Synthetic biology applies engineering principles to create living systems with predictable and useful behaviors from collections of standardized genetic parts. However, living systems unlike mechanical devices inevitably evolve when their DNA sequences accumulate copying errors, often resulting in "broken" cells that no longer function as they were programmed. We are addressing this challenge by better characterizing how engineered cells evolve and using this information to design DNA sequences and host cells that are more robust against unwanted evolution. Also, precisely defining the function, extent, and provenance of genetic parts can be complicated because variation in these parts can arise from engineering or intentional evolution. This work includes: (1) developing "negative design" software to alert researchers to genetically unstable and unintentionally burdensome DNA designs; (2) using experimental evolution and engineering to create "antimutator" variants of host organisms that have lower-than-natural mutation rates; and (3) developing databases and software tools for improved annotation of engineered DNA sequences and common variants of those sequences.
 Current Researchers:
Current Researchers: Dennis Mishler
 Representative Publications
Representative Publications
 Current Funding:
Current Funding: NIH R01;
Past Funding: NSF CAREER, DARPA BRICS, DARPA Insect Allies
Dynamics of Microbial Genome Evolution
The E. coli Long-Term Evolution Experiment (LTEE)
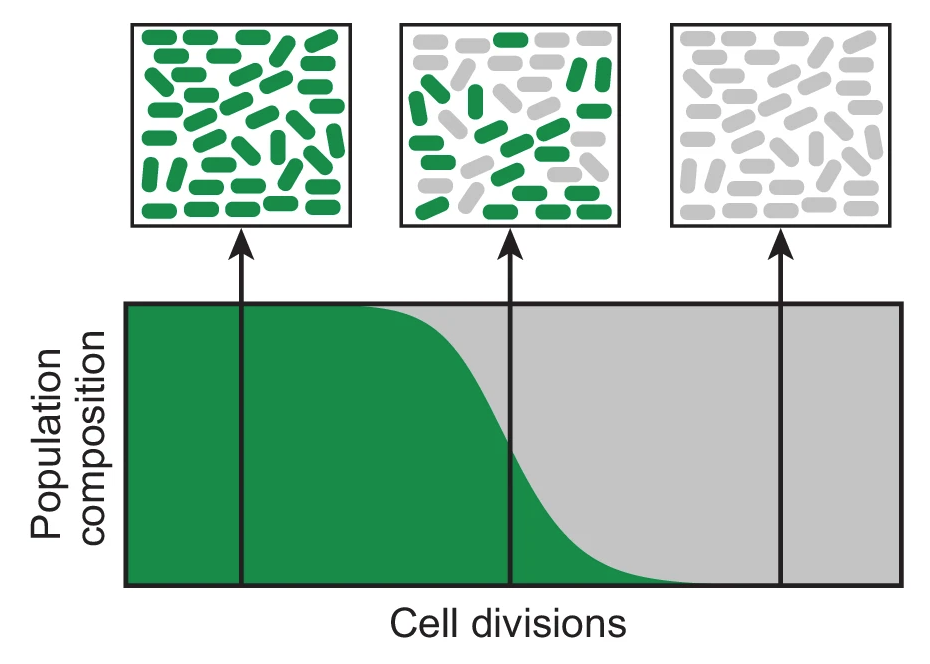
Accumulation of mutations in population Ara-1 of the LTEE over 20,000 generations of evolution
We develop the
breseq computational pipeline for identifying mutations in laboratory-evolved microbial genomes from next-generation sequencing data. We have used this tool to extensively study rates of genome evolution in the >35-year Lenski
long-term evolution experiment (LTEE) with
E. coli. We continue to develop
breseq and related computational pipelines so that it can be used for more additional applications related to strain engineering and medicine. For example, we are interested in how tracking rare variants within populations of microorganisms can anticipate further evolutionary trajectories and how this information might be used to better diagnose and treat disease. We also have other ongoing projects related to understanding the mechanisms and dynamics of bacterial evolution and ecology in the LTEE using other high-throughput and systems biology approaches.
 Current Researchers:
Current Researchers: Alexa Morton, Pranesh Rao, Ira Zibbu, Tyler de Jong
 News: Legendary bacterial evolution experiment enters new era
News: Legendary bacterial evolution experiment enters new era
 Representative Publications
Representative Publications
 Funding:
Funding: NSF LTREB, UT-CNS SPARK, NSF EEID, NASA
Past Funding: NSF K99/R00
Evolution and Engineering of Naturally Transformable Bacteria
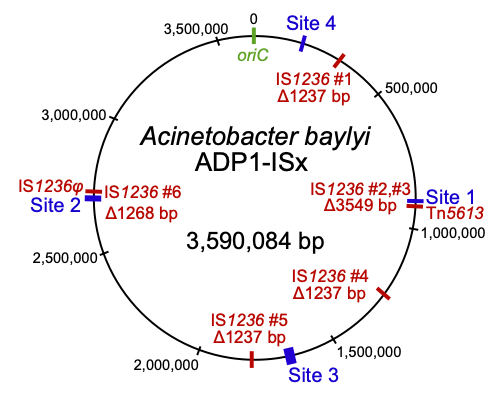
Insertion sequences (red) deleted in the transposon-free strain Acinetobacter baylyi ADP1-ISx
Naturally competent bacteria have expanded evolutionary potential because they can readily acquire new DNA from their environment. We are using experimental evolution of the model organism
Acinetobacter baylyi ADP1 to understand how horizontally acquired genes and mobile genetic elements become domesticated after their incorporation into a new genome and the broader effects of gene acquisition on the rest of the genome. We are also studying the role of chemical specificity in determining the fate of acquired DNA as either nutrition or genetic information. These bacteria also provide an improved platform for studying microbial genome engineering due to the ease of reconstructing mutations and introducing new genes. We are investigating sources of genetic instability in ADP1 and engineering a clean genome version of this strain by deleting transposable elements and prophages. This will promote the use of ADP1 in synthetic biology by reducing rates of mutations that lead to inactivation of introduced genes. Further, we are using ADP1 as a platform to understand the limits to streamlining bacterial genomes and identifying adaptations to overcome the fitness costs of reduced genomes.
 Current Researchers:
Current Researchers: Isaac Gifford, Meghna Vergis
 Representative Publications
Representative Publications
 Funding:
Funding: NSF, UT-CNS SPARK
Past Funding: Welch Foundation, NSF CAREER
Evolution and Ecology of Bartonella Parasites in the Negev Desert
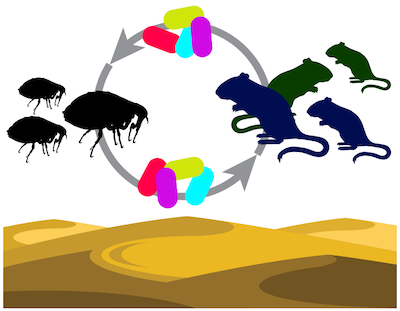
Diverse Bartonella vectored by fleas infect multiple gerbil species in the Negev Desert
Bartonella are diverse blood-borne parasites that can cause zoonotic infections of humans. The
Hawlena Lab studies
Bartonella species that circulate in gerbil populations in the Negev Desert in southwestern Israel. In these natural wildlife communities, multiple species of
Bartonella infect and co-infect multiple host species, making it a dynamic system for studying host-parasite interactions. Our laboratory collaborates on projects characterizing how the genomes of these
Bartonella evolve within and over multiple infection cycles in wild and lab-reared rodents. Our lab is particularly interested in mechanisms for genome evolvability utilized by these bacteria to escape host immune responses, including simple-sequence repeat contingency loci, gene accordions, and recombination during co-infections.
 Current Researchers:
Current Researchers: Isaac Gifford
 Representative Publications
Representative Publications
 Funding:
Funding: BSF
Past Funding: NSF EEID








Barrick Lab > ResearchInterests