Page under construction as of 2/22/2023
Custom Primer Design on Benchling
While there are tools available for automatically designing primers (such as the NCBI Primer BLAST) often specialized PCR applications, such as amplifying fragments for cloning, will necessitate specific primer placements and modifications. In these cases it is important to take extra care to ensure the primers will anneal to the template and amplify well. This page presents guidelines and tools for specifically designing primers from DNA sequences, using the Benchling platform as an example. In short, you will need to: 1) determine where to place your primers within the template sequence to amplify your desired product 2) adjust your primer sequences to ideal annealing temperatures 3) modify the sequences with any necessary adapters and check for inhibitory secondary structures. Designing primers may require trade-offs and revisions. As such it's important to understand the needs of your specific experiment, such as specific locations in the genome, annealing temperatures, and adapter sequences, and keep those in mind while designing your primers. The examples presented here illustrate designing primers to amplify the downstream region of the Acinetobacter baylyi ADP1 ACIAD2049 gene for use in Golden Transformation.1) Determining Primer Placement
Figure 1 Placing a primer downstream of the ACIAD2049 gene.
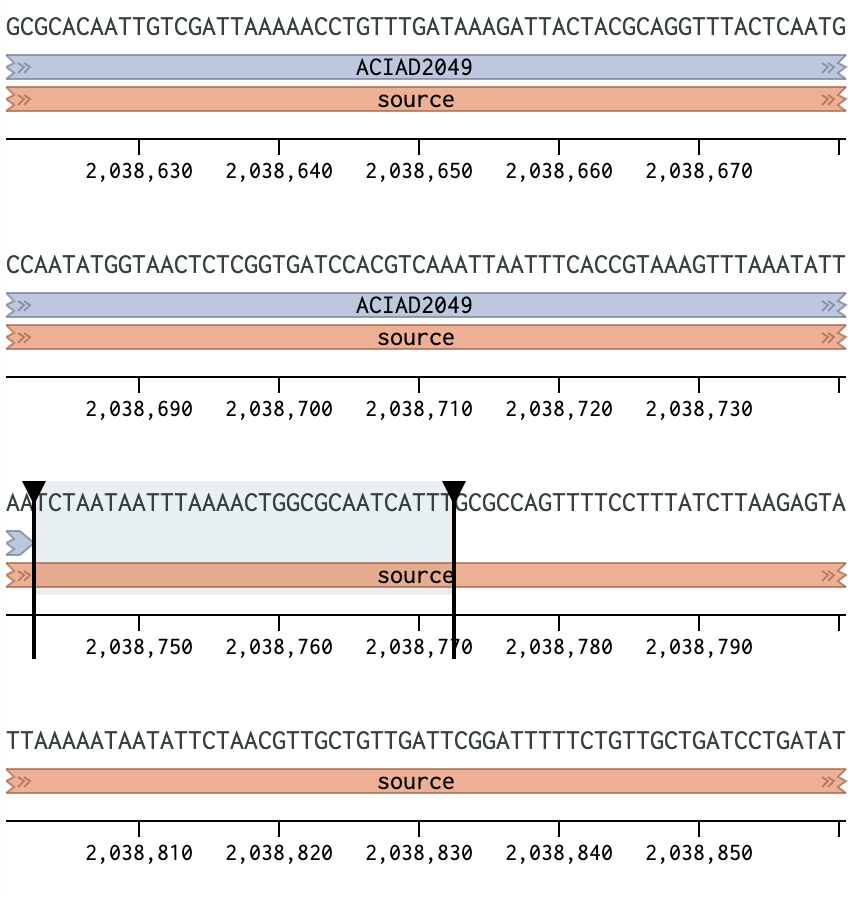
2) Calculating Annealing Temperatures
Calculate the annealing temperature for your PCR reaction using the NEB Tm Calculator. a. Select the polymerase for your PCR reaction from the dropdown menu. b. Copy ~30 bp of the forward and reverse primer sequences into the Primer 1 and Primer 2 boxes. c. Adjust the annealing temperature (top right box) by removing bases from the 3' end of each primer until:- The annealing temperature is in your desired range. This is typically ~57-60°C.
- The individual Tms of the two primers (shown in the boxes on the right) are less than 5° apart.
- Each primer is 18-25 bp long. Shorter primers are preferred because they reduce the possibility of forming secondary structures that may inhibit your reaction.
- Ending a primer with a C or G nucleotide can improve its ability to bind to the template.
Figure 2 Annealing temperature calculations for ACIAD2049 downstream primers.
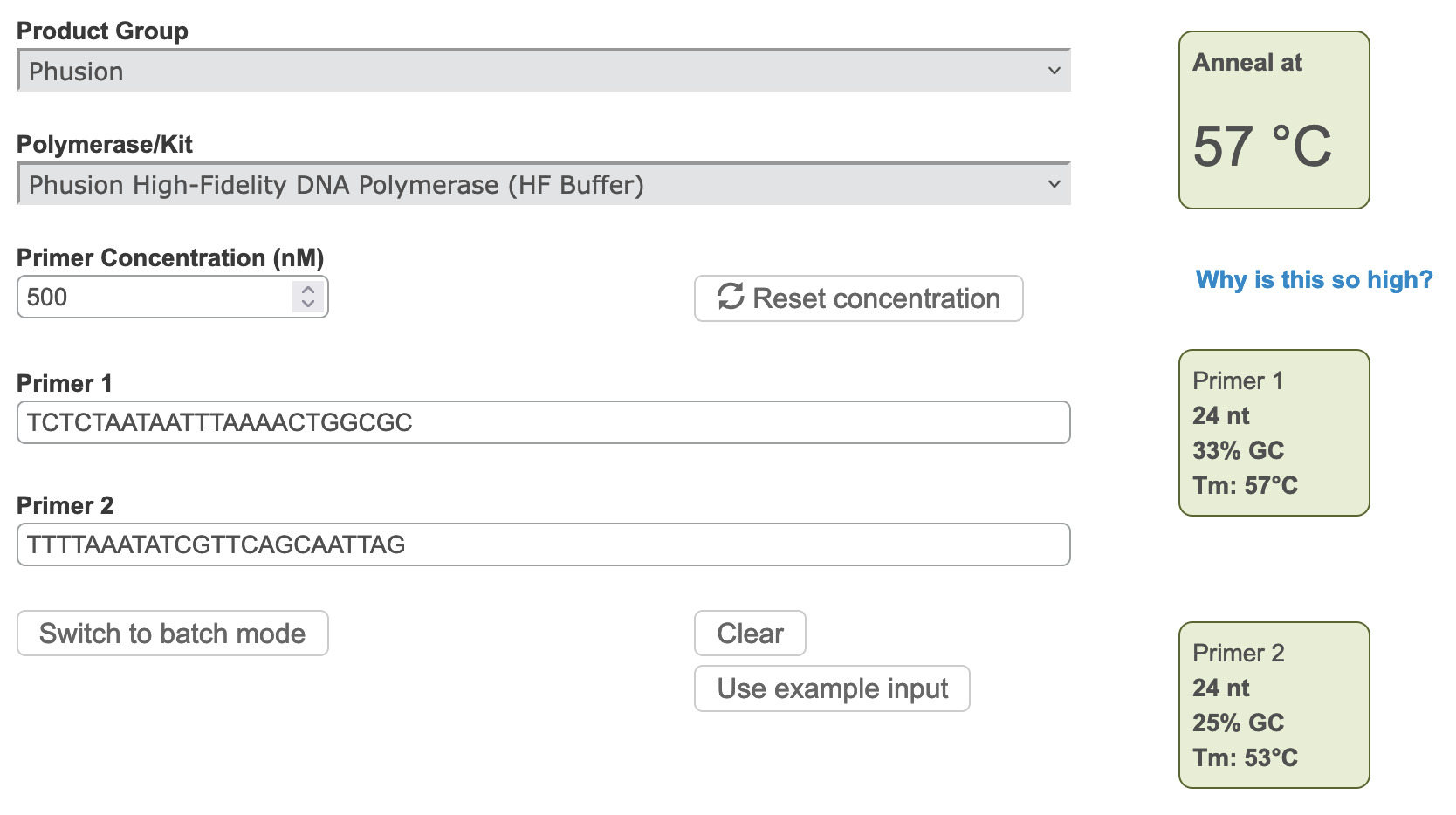
3) Modifying Primer Sequences and Checking Secondary Structures
a. Add any additional modifications or adapters to the end of your sequences. Restriction sequences used for cloning need to be added to the 5' end of the primer sequence. b. Copy and paste each primer sequence into IDT's OligoAnalyzer program. Press "HAIRPIN" to analyze secondary structure formations. Structures with melting temperatures (Tm) below the annealing temperature for your reaction will melt in the thermocycler and not interfere with your reaction (see Figure 3 for example). Note any structures with Tms within a few degrees of your annealing temperature (see Figure 4 for example). c. If problematic structures were detected, modify the primer sequence to remove them. Potential solutions include:- Shortening the primer.
- Relocating the primer binding site to a nearby region of the template DNA (if possible according to the needs of your experiment). This may be especially helpful if your original target site was particularly A/T or G/C rich.
- Choosing a different restriction site or ligation overhang (if possible).
Figure 3 OligoAnalyzer results without problematic secondary structures.
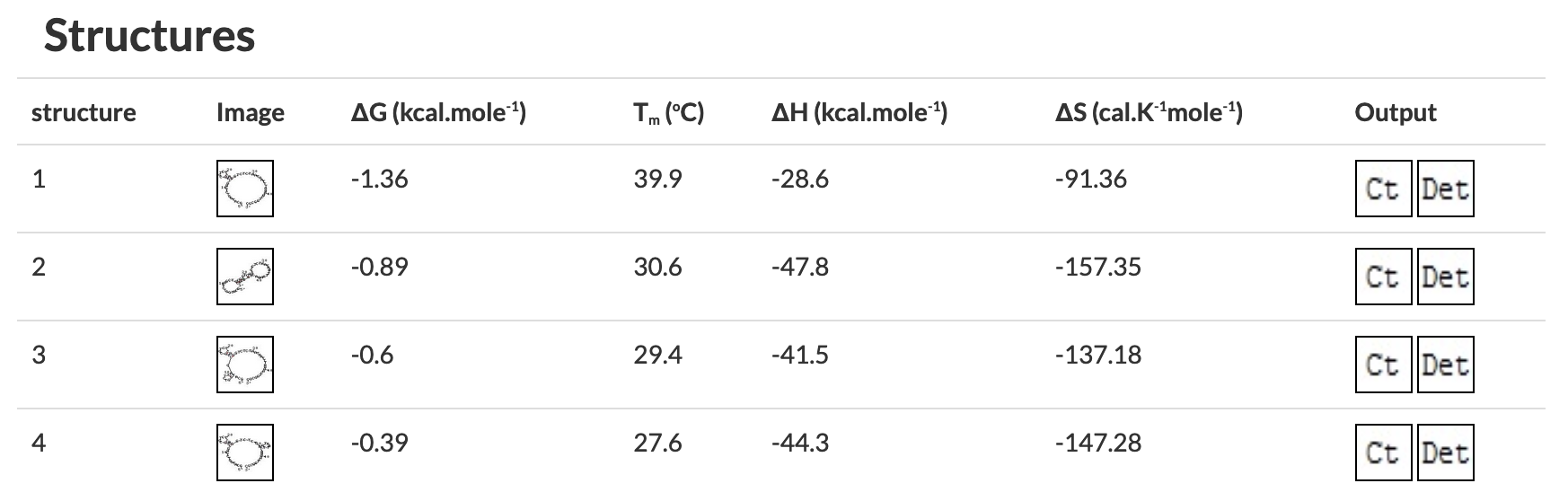
Figure 4 OligoAnalyzer results predicting problematic secondary structures.
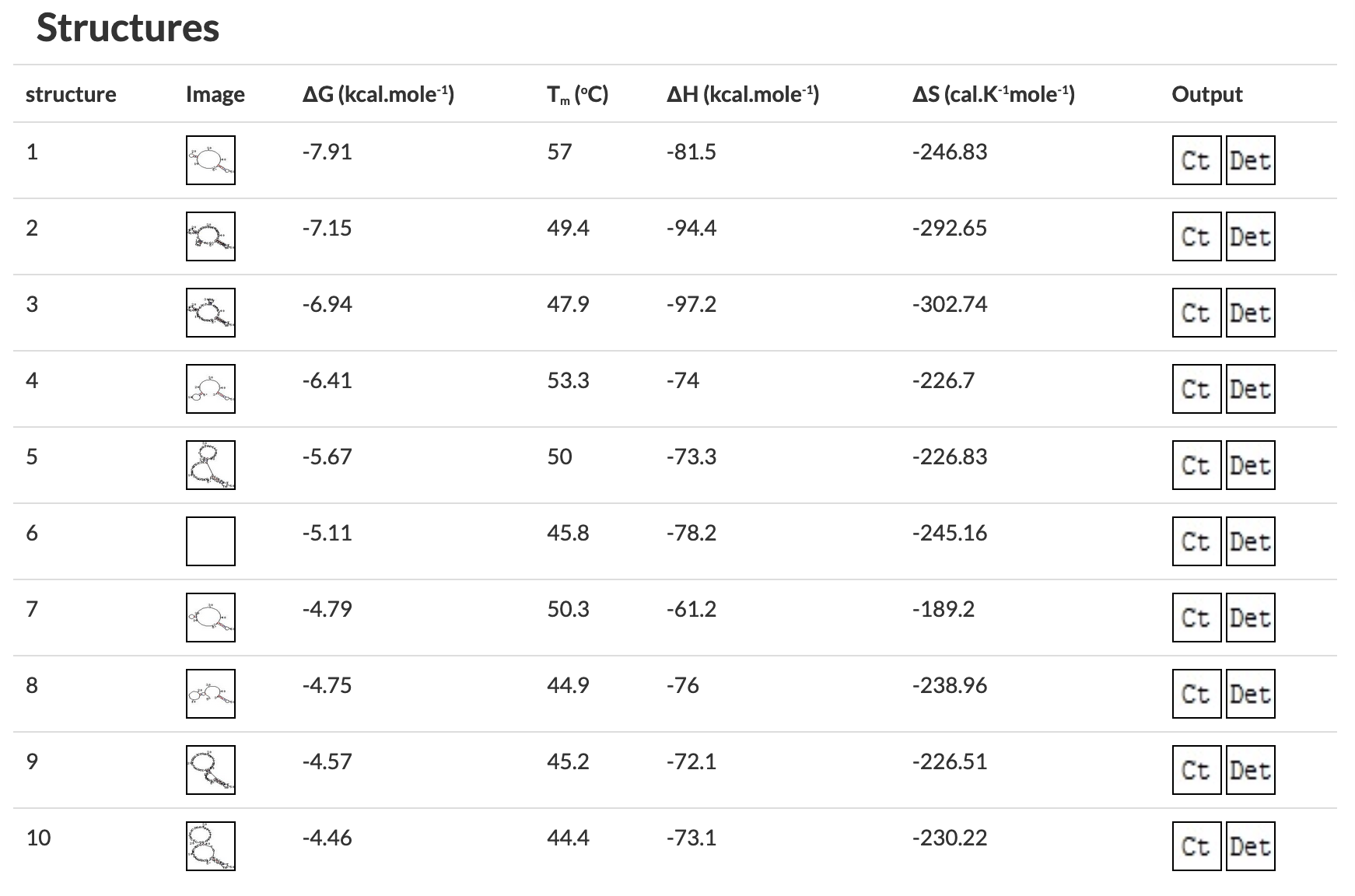
Further Reading
Barrick Lab > ProtocolList > PrimerDesignBenchling
Contributors to this topic  IsaacGifford, CameronRoots, JeffreyBarrick
IsaacGifford, CameronRoots, JeffreyBarrick
Topic revision: r2 - 2023-02-23 - 17:53:00 - Main.IsaacGifford

 Mol Biosciences
Mol Biosciences The LTEE
The LTEE iGEM team
iGEM team NGS course
NGS course