<<Return to qPCR page
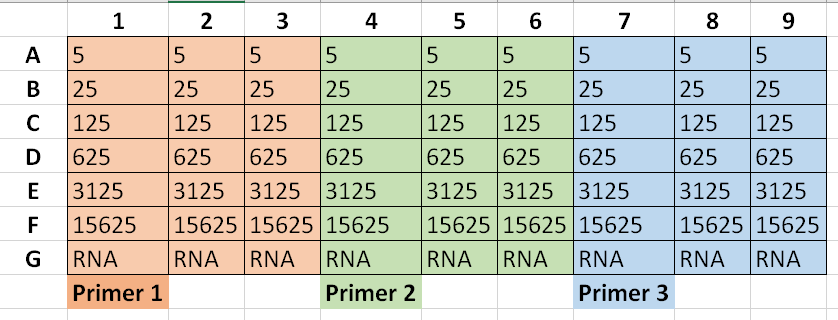 (Numbers are cDNA dilution with "5" representing a 1:5 dilution and so on)
Template Material
(Numbers are cDNA dilution with "5" representing a 1:5 dilution and so on)
Template Material
cDNA Concentration PCR
Goals- Test that primers work
- Verify single products via a melt curve analysis
- Determine what dilution of your cDNA you will use as template in subsequent reactions
- Test RNA for gDNA contamination
 (Numbers are cDNA dilution with "5" representing a 1:5 dilution and so on)
Template Material
(Numbers are cDNA dilution with "5" representing a 1:5 dilution and so on)
Template Material
- The cDNA used for this is a pool of all your cDNA samples i.e. take 1ul of each RT reaction product, pool them, and make dilutions from there. Why? You don't know the difference in expression between your experimental and control sample, or the variation in your biological replicates. If you just ran this plate using cDNA template from one reaction, you might end up choosing conditions suitable for that sample, but not suitable for the others. This limits that error.
- The RNA sample used to test for gDNA contamination is a pool of all your RNA samples (just like you did for the cDNA pool in the step above). This pool is then diluted to the same extent as the 1:25 cDNA dilution i.e. take Xng of your pooled RNA (x = ng you inputted into RT), make up to 20ul (because your initial reverse transcription was 20ul), and then make a 1:25 dilution.
- I would recommend testing the cDNA concentration using representative primers for each condition. For example, pick one reference gene, one target gene that you might expect to be highly expressed, and one target gene that you expect to not be expressed as much. I like to at least test 3 primers in case there is an outlier. This is important because you want to make sure that all your test conditions fall within the dynamic range.
- PCR products that produce a single peak in your melt curve analysis
- A dilution of cDNA that produces CT values of between 13-30 cycles (If your CTs are less than or greater than this, the assay can still work, but depending on machine, standard deviations can get a bit noisy if amplification happens too early or late.)
- No contamination in RNA, or contamination that is >5 cycles later than the signal in your experimental sample. For primers that are 100% efficient, each cycle represents a doubling of product. Therefore a difference of 7 cycles would be a difference in expression of 2^7 or 128 fold.
Analysis
There isn't any real analysis for this step. The goal is to pick a cDNA dilution to use in subsequent steps. Follow the "what you are looking for" guidelines and pick the dilution that best matches those criteria! Once you've diluted all of your cDNA samples store them at 4°C to avoid complications with freeze/thaw <<Return to qPCR page Barrick Lab > ProtocolList > QPCR > CDNAConcentrationqPCR