
Difference: MEGAWHOP (9 vs. 10)
Revision 102017-06-13 - GabrielSuarez
Megaprimer whole plasmid cloning | |||||||||
| Changed: | |||||||||
| < < | aka MEGAWHOP cloning | ||||||||
| > > | aka MEGAWHOP cloning aka Overlap Extension PCR cloning | ||||||||
| Deleted: | |||||||||
| < < | aka Overlap Extension PCR cloning | ||||||||
Adapted from Bryksin AV, Matsumura I. 2010. Overlap extension PCR Cloning: a simple and reliable way to create recombinant plasmids. Biotechniques.48(6):463-5.
PurposeTo insert a DNA sequence into a plasmid without restriction enzymes.Experimental Steps
| |||||||||
| Changed: | |||||||||
| < < | | ||||||||
| > > | |||||||||
| Deleted: | |||||||||
| < < | |||||||||
Designing Primers | |||||||||
| Changed: | |||||||||
| < < | 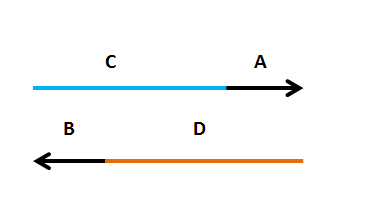 | ||||||||
| > > |  | ||||||||
Primers need to have two components
PCR Insert | |||||||||
| Changed: | |||||||||
| < < | Use stardard 25ul Phusion (or other high fidelity polymerase) protocol | ||||||||
| > > | Use standard 25ul Phusion (or other high fidelity polymerase) protocol | ||||||||
| |||||||||
| Changed: | |||||||||
| < < |
| ||||||||
| > > |
| ||||||||
| |||||||||
| Changed: | |||||||||
| < < |
| ||||||||
| > > |
| ||||||||
| |||||||||
| Changed: | |||||||||
| < < | Purify PCR products. This can either be done through a gel extraction or by adding Dpn1 directly to the Phusion reaction mixture after PCR, digesting at 37°C for 1 hour, and then doing a standard PCR clean up. Dpn1 works efficiently in a Phusion reaction mixture. | ||||||||
| > > | Purify PCR products. This can either be done through a gel extraction or by adding Dpn1 directly to the Phusion reaction mixture after PCR, digesting at 37°C for 1 hour, and then doing a standard PCR clean up. Dpn1 works efficiently in a Phusion reaction mixture. | ||||||||
PCR Recombinant Plasmid | |||||||||
| Changed: | |||||||||
| < < | Use modified 10ul Phusion (or other high fidelity polymerase) protocol | ||||||||
| > > | Use modified 10ul Phusion (or other high fidelity polymerase) protocol | ||||||||
| |||||||||
| Changed: | |||||||||
| < < |
| ||||||||
| > > |
| ||||||||
| |||||||||
| Changed: | |||||||||
| < < |
| ||||||||
| > > |
| ||||||||
| |||||||||
| Changed: | |||||||||
| < < | Adjust Elongation time for the length of the entire plasmid (90 seconds per kb). | ||||||||
| > > | Adjust Elongation time for the length of the entire plasmid (90 seconds per kb). | ||||||||
| OR | |||||||||
| Changed: | |||||||||
| < < | |||||||||
| > > | Use modified 25ul Phusion protocol | ||||||||
| Deleted: | |||||||||
| < < | Use modified 25ul Phusion protocol | ||||||||
| |||||||||
| Changed: | |||||||||
| < < |
| ||||||||
| > > |
| ||||||||
| |||||||||
| Changed: | |||||||||
| < < | 68°C 5min+98°C 3min+(98°C 30s+68°C 30s+72°C X min)*30 +72°C 10min | ||||||||
| > > | 68°C 5min+98°C 3min+(98°C 30s+68°C 30s+72°C X min)*30 +72°C 10min Adjust Elongation time for the length of the entire plasmid (30 seconds per kb). | ||||||||
| Deleted: | |||||||||
| < < | Adjust Elongation time for the length of the entire plasmid (30 seconds per kb). | ||||||||
| Changed: | |||||||||
| < < | DigestOnce the reaction is complete, digest the Recombinant Plasmid PCR product with 0.5 ul Dpn1 at 37°C for one and a half hours to remove parental DNA. DpnI Digest | ||||||||
| > > | DigestOnce the reaction is complete, digest the Recombinant Plasmid PCR product with 0.5 ul Dpn1 at 37°C for one and a half hours to remove parental DNA. DpnI Digest | ||||||||
TransformHeatshock 5ul of the Dpn1-digested MEGAWHOP reaction mixture into 25ul of chemically competent E coli. | |||||||||
| Deleted: | |||||||||
| < < | |||||||||
ExampleChange the promoter for sgRNA using MegaWHOP. Template sequence: https://benchling.com/s/g4S95i24 | |||||||||
| Changed: | |||||||||
| < < | Primers: | ||||||||
| > > | Primers: | ||||||||
| |||||||||
| Changed: | |||||||||
| < < |
| ||||||||
| > > |
| ||||||||
| UPCASE LETTERS: a region that amplifies the insert (A(or B) | |||||||||
| Changed: | |||||||||
| < < | lower case letters: a region that targets the new plasmid (C(or D) | ||||||||
| > > | lower case letters: a region that targets the new plasmid (C(or D) | ||||||||
| Changed: | |||||||||
| < < | *PCR MEGA_primer: | ||||||||
| > > | *PCR MEGA_primer: 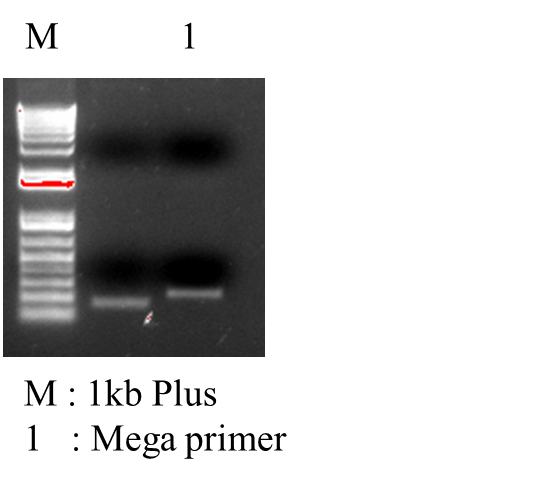 | ||||||||
| Deleted: | |||||||||
| < < |  | ||||||||
| Changed: | |||||||||
| < < | *PCR Recombinant Plasmid: | ||||||||
| > > | *PCR Recombinant Plasmid: 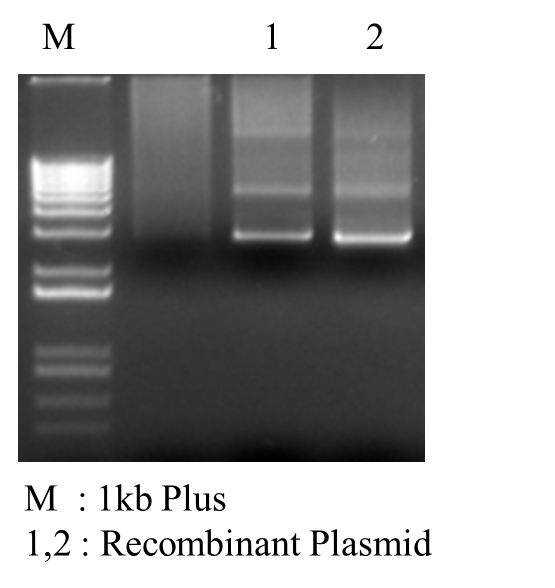 | ||||||||
| Deleted: | |||||||||
| < < | 
| ||||||||
| |||||||||
View topic | History: r12 < r11 < r10 < r9 | More topic actions...