Protocol for harvesting _pfu-sso7d (aka Phusion) polymerase
This protocol is for expressing and purifying the high fidelity
pfu-sso7d polymerase [1] from
E. coli. This protein is sold as Phusion polymerase by
New England Biolabs. This
pfu variant has the
sso7d processivity-enhancing domain attached that increases its speed and processivity. It generates blunt-end DNA products and typically you use higher annealing temperatures than when using
taq.
See the
NEB website for a description of other key enzyme characteristics.
Expression plasmid sequence:
6his-pfu-sso7d-pET28.gbk
Materials needed:
- Glycerol stock of EQ458 E. coli cells
The strain used is named EQ458. It is located in common species box; this is a Rosetta 2 (DE3) E. coli strain containing 6his-pfu-sso7d-pET28 plasmid. The plasmid is KanR and the strain itself is CamR. The frozen stock is overnight growth of a single colony.
- LB medium. Link to LB recipe
- Chloramphenicol stock. Link to Cam recipe
- Kanamycin stock. Link to Kan recipe
- Refrigerated centrifuge.
- Spectrophotometer and cuvettes.
- French press. Georgiou lab, MBB, 3.310, ask before using.
- IPTG, 100 mM stock. Dissolve 2.38 g IPTG in 100 mL deionized water. Filter sterilize and store at 20°C.
- Disposable plastic columns. ThermoSci, cat #29922
- Ni-NTA agarose resin. Qiagen, cat #30210, 25 ml
- Slide-A-Lyzer, 10k dialysis cassette G2. ThermoSci, cat# 87730
Lysis Buffer:
- 50 mM NaH2PO4
- 300 mM NaCl
- 10mM Imidazole
- Adjust pH to 8.0 using NaOH
Wash Buffer:
- 50 mM NaH2PO4
- 300 mM NaCl
- 40mM Imidazole
- Adjust pH to 8.0 using NaOH
Elution Buffer:
- 50 mM NaH2PO4
- 300 mM NaCl
- 250 mM Imidazole
- Adjust pH to 8.0 using NaOH
Polymerase storage buffer: Make 3-4 Liters
- 50% Glycerol
- 100 mM Tris/HCl pH 8.0
- 0.2 mM EDTA
- 0.2% NP-40; nonionic detergent
- 0.2% Tween20
- 2 mM DTT (add immediately before use)
IMPORTANT: Add fresh DTT immediately before use by freshly dissolving it from powder or from a 1 M stock stored at 20°C. We have observed rapid loss of function of enzyme when it is diluted in old storage buffer that has been stored at room temperature.
Protein Expression
Scaled for 2 x 500 mL cultures
Day 1: Revive and Isolate Colony
- Streak LB plate supplemented with Kan and Cam from frozen stock of EQ458. Growth plate overnight at 37°C.
Day 2: Precondition
- Select single colony from O/N streak plate and inoculate 1.5 mL of LB broth supplemented with Kan and Cam. Grow overnight at 37 C shaking at 250 rpm.
Day 1: Induce
- Use 500 µL of overnight culture to inoculate 500 mL of supplemented LB broth (in 2 L flask), grow as before for ~ 3-4 hours until an OD600 of between 0.4 and 0.6 is reached.
- Induce the cultures to express proteins by adding IPTG at a final concentration of 0.5 mM (2.5 mL per 500 mL) followed by overnight growth at 18 C, 250 rpm.
Day 0: Harvest
- Collect cells by centrifugation. Conditions as follows: 4°C, at 10,000 x g for 15 mins.
- Resuspend each cell pellet with 3 mL of lysis buffer and combine tubes together, mix well using pipette.
- French press; use the full cell holding (10 mL - 35 mL) and 1500 psi pressure.
- Collect and reintroduce into french press 1x.
- Heat denature at 70°C for about 15 mins.
- Spin down, 10,000 x g for 30 mins.
- Syringe filter the supernatant (0.22 µm filter).
- Proceed to IMAC purifications.
Immobilized metal ion affinity chromatography (IMAC) purification
Note: Save portions at each step for protein gel
- Prepare a 1 mL Ni-NTA resin column.
- Saturate column with 5x the column volume (so 5 mL) of lysis buffer. Repeat this step twice.
- Bind with 1:1 lysis buffer:SN sample.
- Wash with 1x column volume of lysis buffer.
- Wash with 5x (column volume) of wash buffer.
- Elute with 3 mL of elution buffer and collect all 3 mL of elution.
Dialysis:
- Place dialysis cassette into storage buffer for 2 mins.
- Remove top and load dialysis cassette with enzyme sample using a pipette or syringe.
- Squeeze the membrane to remove excess air.
- Replace top and place in beaker with 500 mL or 1 L of storage buffer. This should be done in the cold room on a stir plate.
- Allow to sit for 2 to 4 hours.
- Remove cassette and place in beaker with fresh storage buffer. Allow to sit overnight.
- If cassette has swollen, use syringe to remove some of the sample.
- Open top of cassette and remove sample.
- Store at 20°C.
Assay purified phusion polymerase activity by PCR
* IMPORTANT: You must use commercial Phusion buffer (
NEB Cat #B0518S) for your reactions. It is a proprietary formulation that gives MUCH better enzyme performance.
- Template for this assay is the 6his-pfu-sso7d-pET28 plasmid encoding the phusion polymerase.
- To estimate the activity of your purified Phusion, create a dilution series of purified polymerase in water ranging from 1:200 to 1:10, and compare to NEB's stock.
- NEB stock is viscous; for an accurate comparison to the purified Phusion, ensure you are pipetting sufficient volumes to maintain accuracy.
Primer sequences (position with reference to sequence in file above):
| Name |
Position |
Tm |
5' - 3' |
Amplicon size w/ R1 (bp) |
| Phusion F1 |
672-695 |
66.4 |
agttccataggatggcaagatcc |
4044 |
| Phusion F2 |
2748-2770 |
66.5 |
tgataccgatgaaacgagagagg |
1968 |
| Phusion F3 |
3759-3779 |
66.4 |
gagctgtcttcggtatcgtcg |
957 |
| Phusion F4 |
4169-4190 |
66.7 |
aacattagtgcaggcagcttcc |
547 |
| Phusion R1 |
4694-4716 |
67.8 |
cctaatgcaggagtcgcataagg |
NA |
Protocol:
| Reagent |
Volume/ul |
| Water |
12.4 |
| dNTP (10mM) |
0.4 |
| 5x buffer |
4 |
| Primer F (10uM) |
1 |
| Primer R (10uM) |
1 |
| Template (2ng/ul) |
1 |
| Diluted Phusion |
0.2 |
Conditions (Denaturation-Annealing-Extension repeated 30x):
| Cycle |
Temperature |
Duration (secs) |
| Initial denaturation |
98 |
30s |
| Denaturation |
98 |
10s |
| Annealing |
69 |
20s |
| Extension |
72 |
30s / kbp |
| Final extension |
72 |
5m |
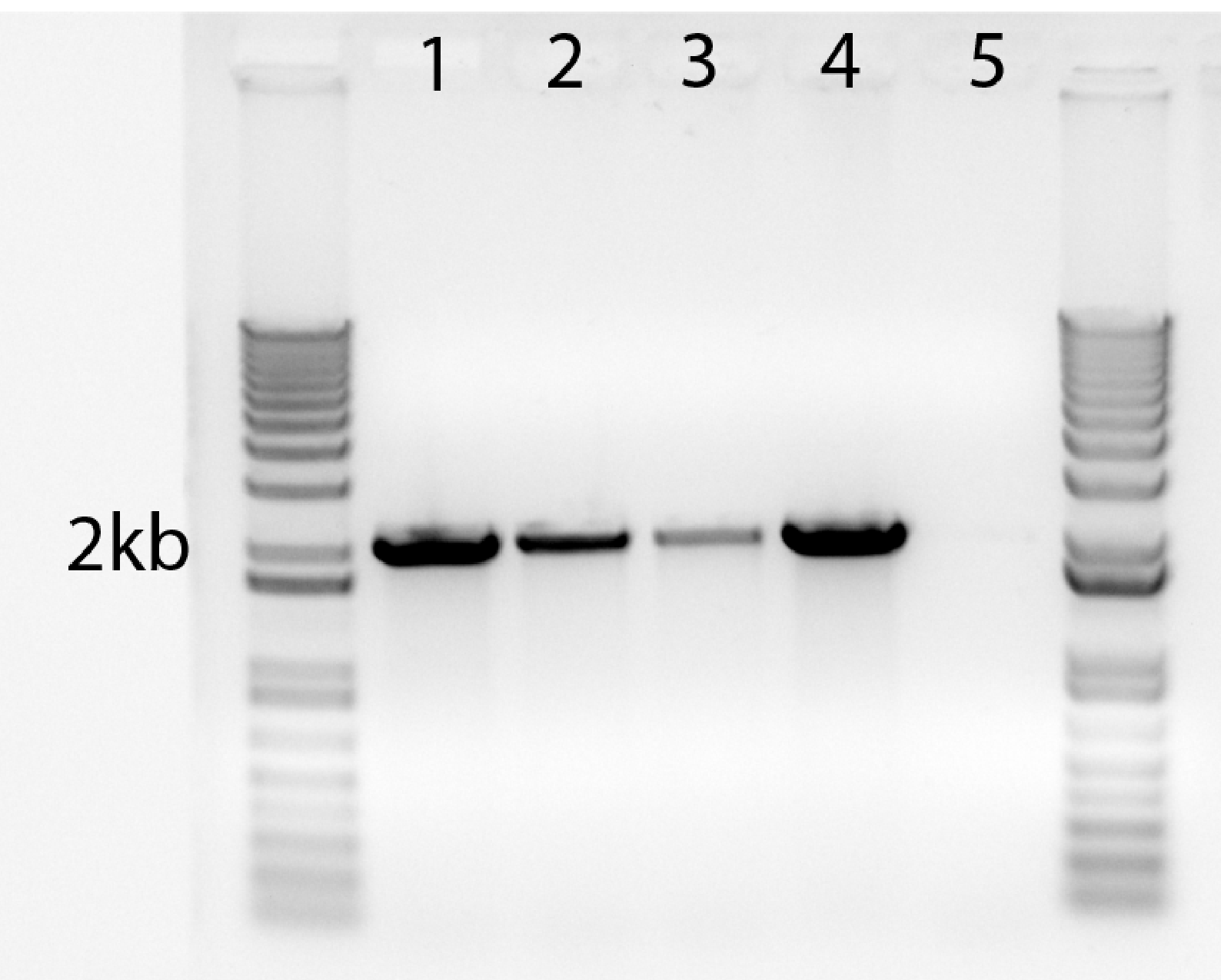
Sample results for 2kbp amplicon:
| Lane |
1 |
2 |
3 |
4 |
5 |
| Phusion dilution |
25 |
50 |
100 |
NEB (neat) |
H20 |
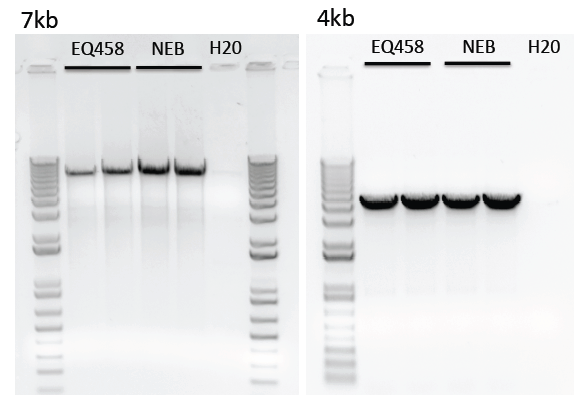
Limitations of purified polymerase
To date, we have noted reduced processitivity of longer transcripts (>4kb) using our purified polymerase compared to the commercial variant.
References
1. Wang, Y., Prosen, D. E., Mei, L., Sullivan, J. C., Finney, M., Vander Horn, P. B. (2004) A novel strategy to engineer DNA polymerases for enhanced processivity and improved performance in vitro.
Nucleic Acids Res. 32: 11971207.
«PubMedCentral»
Barrick Lab > ProtocolList > ProtocolsReagentRecipes > ProtocolsReagentsPfuSso7d
 JeffreyBarrick, CraigBarnhart, SimonDAlton
JeffreyBarrick, CraigBarnhart, SimonDAlton


 Mol Biosciences
Mol Biosciences The LTEE
The LTEE iGEM team
iGEM team NGS course
NGS course