Gibson Assembly
Background and Design
Gibson Cloning is a technique of DNA construct assembly that allows one to join multiple linear segments into either one large linear segment or, if the segments contain the appropriate components and overlaps, an intact plasmid. This protocol follows the one-step ISO assembly of overlapping dsDNA protocol. In order to assemble segments of DNA via Gibson Cloning, they must contain at least 40bp of homology to the segment they are being joined to. For example, if one was to make a construct that was Seg1-Seg2-Seg3 from individual PCRs of Seg1, Seg2, and Seg3, the 3' end of Seg1 would need 40bp of homology to the 5' end of Seg2 and Seg3 would require it 5' end to have 40bp of homology to Seg2. For constructing a plasmid, use a linearized vector backbone as one of your segments.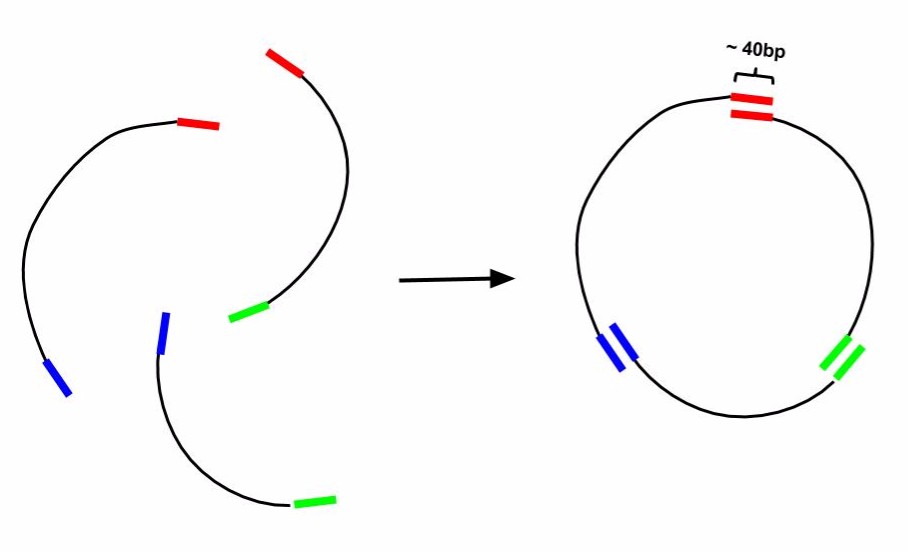
Supplies
If master mix aliquots are available, and less than 1 year old:- 15uL aliquots of Gibson master mix, located on the top shelf of the -20°C in Welch. Tubes are in both a 96 well holder, and a 50mL tube.
- Pure water
- Positive control: Positive Control DNA Mix (see below)
Primer Design Using Gibson
For a thorough discussion on the construction of primers for use in Gibson Assembly, please see the following publication: http://www.ncbi.nlm.nih.gov/pubmed/21601685.Protocol
- Use PCR to produce the DNA segments needed for assembling the new construct. Confirm the success of each PCR by running 5uL of the reaction on an agarose gel.
- Mix 10ng-100ng of each of your DNA segments together (such that their ratios are equimolar) into a 5uL total volume. Therefore, the length of each fragment, and the concentration of the fragments must be taken into account.
- Add that 5uL of DNA to the Gibson Master Mix, mix well by pipetting.
- Incubate the reaction at 50C for 1 hour.
- Dilute the reaction 1:5 in sterile, nuclease free water. Use this diluted sample for PCR (if linear) or transformation (if a plasmid).
- Run an aliquot of your final reaction on a gel to verify the presence of your construct. It may be used to do make a 'no incubation' control to determine if the chemistry of the reaction happened.
Expected Results
The positive control should yield ampicillin resistant colonies when transformed into E. coli.Positive Control Construction and Usage
The Positive Control DNA Mix for Gibson Assembly consists of a two-piece assembly of pUC19. It is designed such that 5uL of the Positive Control DNA Mix is to be added to 15uL of Gibson Assembly Master Mix along side experimental reactions. Both pUC19 segments are between 1.3kb and 1.4kb in size. To construct the positive control reaction mix:- PCR amplify the two pUC19 fragments - fragment 1 (F1) and fragment 2 (F2).
- Use primers pUC19 F1 Gib FW (5'-CTCTTACTGTCATGCCATCCGTAAGATGCTTTTCTGTGACTGG-3') and pUC19 F1 Gib RV (5'-ACATGTTCTTTCCTGCGTTATCCCCTGATTCTGTGGATAA-3') to create F1 and pUC19 F2 Gib FW (5'-TTATCCACAGAATCAGGGGATAACGCAGGAAAGAACATGT-3') and pUC19 F2 Gib RV (5'-CCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAG-3').
- Use the following reaction mixture:
- 32.7uL dH2O
- 10uL 5X HF Buffer
- 1uL 10uM dNTP
- 2.5uL 10uM FW primer
- 2.5uL 10uM RV primer
- 1uL ~3ng/uL pUC19 template
- 0.3uL phusion polymerase
- Use the following PCR program for both F1 and F2 PCRs:
- 98C - 0:30
- 98C - 0:10
- 58C - 0:30
- 72C - 0:45
- Go to step 2, 25X
- 72C - 10:00
- After the reaction completes, add 1uL of Dpn1 (20,000 U/mL) to each reaction, briefly vortex and spin down the reactions, and then incubate in the PCR machine at 37C for 30 minutes. To test the effectiveness of your Dpn1, construct the above PCR reaction without phusion polymerase, digest and purify it in parallel with the F1 and F2 PCRs, and transform it along side your other reactions. Any ampicillan-resistant transformants produced from from transforming this negative reaction indicate your Dpn1 digest was ineffective at digesting the pUC19 template. The solution to this is to use fresh Dpn1 in your digest, or gel extract F1 and F2 instead of using a Dpn1 digest and column purification to remove pUC19 template.
- Test the success of the PCR by gel electrophoresis and purify the reaction using a PCR cleanup kit, and elute with buffer or water without EDTA. Quantify the concentration of F1 and F2 purifications.
- Combine F1 and F2 PCR products to a final concentration of 2.8ng/uL of each fragment. Dilute with the elution buffer used in the PCR purification if needed. This final mixture is the Positive Control DNA Mix.
| I | Attachment | History | Action | Size | Date | Who | Comment |
|---|---|---|---|---|---|---|---|
| |
Gibson_Assembly_pic.jpg | r1 | manage | 29.5 K | 2016-12-16 - 15:17 | SimonDOelsnitz |
Barrick Lab > ProtocolList > ProtocolsGibsonCloning
Contributors to this topic  NeilGottel, BrianRenda, SimonDOelsnitz, DaciaLeon, MattMcGuffie, CameronRoots, SimonDAlton, JeffreyBarrick
NeilGottel, BrianRenda, SimonDOelsnitz, DaciaLeon, MattMcGuffie, CameronRoots, SimonDAlton, JeffreyBarrick
Topic revision: r10 - 2016-12-16 - 15:20:56 - Main.SimonDOelsnitz



 Mol Biosciences
Mol Biosciences The LTEE
The LTEE iGEM team
iGEM team NGS course
NGS course