Back to Golden Gate Protocols
2. Mix samples well by pipetting, then run the reaction on the thermocycler under the following conditions:
3. Transform 2 μL assembly reaction into Electrocompetent or Chemically Competent cells and plate on LB + Cam
Back to Golden Gate Protocols
Part Plasmid Assembly
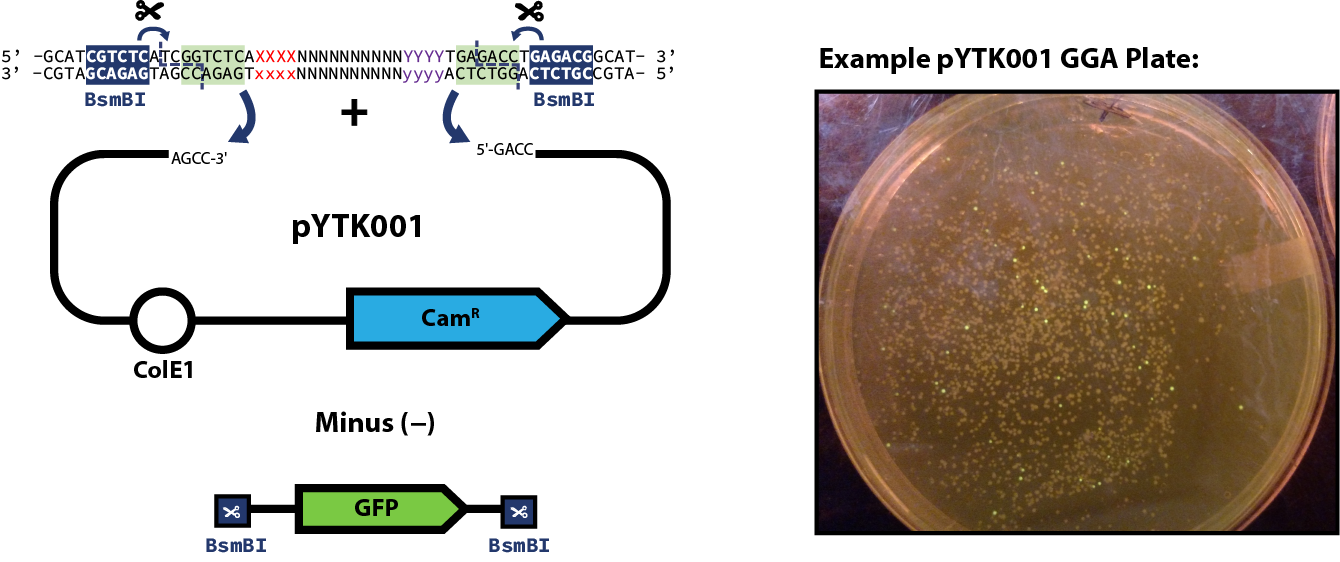
Once you have designed your part and either amplified with PCR or ordered the desired gBlock (as described here) you can proceed to the assembly step of the part plasmid itself. The example reaction below shows pYTK001 used as the entry vector for the reaction; however, this can be substituted for any other entry vector with requisite BsmBI cut sites. Once built, part plasmids can be assembled into transcriptional units.Assembly Reaction
Access the old non-kit golden gate assembly protocols here 1. Set up the following reaction mix:- 10 fmol pYTK-001 plasmid = 17.7ng [try to keep volume to 1-2μL]
- 20 fmol of insert DNA = 650 * insert length * 20x10^-6 = X ng [try to keep volume to less than 10 μL]
- 2 μL of 10× T4 DNA ligase buffer (Promega)
- 1 μL of BsmBI
- 1 μL of T4 DNA ligase
- x μL water up to 20 μL total
| Step | Temperature | Time |
|---|---|---|
| 1 | 42°C | 1.5 min |
| 2 | 16°C | 3 min |
| Cycles 1-2: | Repeat 25x | |
| 3 | 50°C | 5 min |
| 4 | 80°C | 10 min |
Expected Results

- (If using pYTK001 as your entry vector) Multiple non-fluorescent colonies
- Be sure to screen colonies for GFP expression on a blue light transilluminator, pick colonies that do not fluoresce!
- Cells containing properly assembled plasmids (confirmed through PCR and Sanger Sequencing)
Back to Golden Gate Protocols
| I | Attachment | History | Action | Size | Date | Who | Comment |
|---|---|---|---|---|---|---|---|
| |
Part_Plasmid_Assembly.png | r2 r1 | manage | 421.2 K | 2021-07-15 - 22:52 | KateElston |
This topic: Lab > BroadHostRangeToolkit > ProtocolsBTKMakeANewPartPlasmid
Topic revision: r12 - 2021-11-03 - KateElston

